Abstract
Background: Swab screening for vancomycin-resistant Enterococcus (VRE) is used to identify VRE carriers and thus prevent transmission of this organism. We tested whether screening results could have unintended benefits in predicting the vancomycin susceptibility of subsequent Enterococcus infections experienced by these patients.
Methods: We conducted a retrospective cohort study of all patients admitted to a large tertiary care hospital between 2010 and 2015 who underwent swab screening for VRE before experiencing an Enterococcus sterile-site infection. We derived test characteristics using the screening result as the test variable predicting the presence or absence of vancomycin resistance in the sterile-site Enterococcus species.
Results: Culture results for sterile-site samples were positive for Enterococcus for 619 patients during the study period. Of these, 488 (79%) had previously undergone VRE screening. A total of 19 (4%) of the 488 screening results were positive for VRE, and 10 (2%) of the sterile-site Enterococcus isolates were resistant to vancomycin. The overall specificity of VRE swabs was 97% (95% confidence interval [CI] 96%-99%), sensitivity was 70% (95% CI 35%-93%), the positive likelihood ratio was 28 (95% CI 14-56), and the negative likelihood ratio was 0.31 (95% CI 0.12-0.79). The post-test probability of vancomycin resistance in a sterile-site culture, given a positive result with VRE screening swab, was 37% using study data and 83% using published aggregate prevalence data for VRE in the United States.
Interpretation: Prior VRE screening swab results represent a useful tool for predicting vancomycin resistance in sterile-site Enterococcus infections. Patients with a positive result for VRE screening swab and Enterococcus identified by culture of sterile-site samples should receive empiric treatment with linezolid or daptomycin until sensitivity results are available.
Enterococcus is among the top causes of bloodstream infection, infective endocarditis, intra-abdominal sepsis and surgical-site infection.1-5 Colonization with vancomycin-resistant Enterococcus (VRE) is associated with antibiotic use, residence in a long-term care facility, renal disease, malignant disease, organ transplant, critical illness, immunocompromise, catheter use and HIV infection.6-9 Few antibiotic agents are effective against VRE, and they are not often used in initial empiric treatment regimens.10,11 Among patients with sterile-site infections caused by Enterococcus, vancomycin resistance has been associated with higher mortality.10 A recent study showed that a delay of more than 2 days in receiving effective therapy for VRE infection was associated with a 3-fold increase in 30-day mortality.12
The timely identification of antibiotic resistance is challenging. Traditionally, Gram staining results are available early, whereas identification of species has required 2-3 days and determination of susceptibilities longer still. New, rapid diagnostic tools allow for species identification up to 30 hours earlier, which has increased the importance of species-based empiric therapy.13 As a result, ways are needed to predict the interim likelihood of vancomycin resistance in known Enterococcus isolates. The use of swab screening for VRE is attractive because such screening is frequently performed upon admission to hospital. In cases of infection with methicillin-resistant Staphylococcus aureus, results obtained with screening swabs have had high specificity for predicting methicillin resistance in subsequent S. aureus infections.14 Previous studies have evaluated the predictive value of VRE screening swabs in forecasting the prevalence of VRE among all causes of bacterial infection.15-19
In this study, we sought to investigate the utility of results obtained with VRE screening swabs in predicting vancomycin resistance among subsequent sterile-site infections with Enterococcus. We hypothesized that among patients with sterile-site Enterococcus infections, prior positive results with VRE screening swabs would reliably predict vancomycin resistance, but prior negative results would not reliably rule out vancomycin resistance.
Methods
Design and patient selection
We performed a retrospective cohort study involving all patients at Sunnybrook Health Sciences Centre, a large tertiary care centre in Toronto, Ontario, who had a sterile-site isolate of Enterococcus sp. and prior swab screening for VRE between Apr. 1, 2010, and Dec. 31, 2015. We defined sterile-site isolates as isolates from blood, tissue biopsy specimens and cerebrospinal, pleural, peritoneal or synovial fluid. We included patients who were older than 1 year and had samples taken for culture as outpatients, in the emergency department or during hospital admission (excluding long-term care wards). Each patient was included in the study only once, with the most recent Enterococcus isolate and the most recent VRE screening result being used for the study. We excluded patients whose VRE screening was performed after the sample yielding a sterile-site isolate was obtained.
Data sources
We derived data from the Stewardship Program Integrated Resource Information Technology database, which is securely and confidentially autopopulated with pharmacy and microbiology data, as well as demographic characteristics, for Sunnybrook patients (inception April 2010).20
VRE screening protocols and methods
Screening for VRE, via rectal swabbing, was performed on the first day of admission under the following circumstances: known previous colonization, infection or contact with a resistant organism; previous receipt of health care in a high-risk area outside of Canada; admission to a health care facility within the preceding year; receipt of home health care or hemodialysis; residence in a shelter or other high-risk setting; or inability to answer screening questions. Surveillance swabs were also obtained during transfer to a critical care unit or in the context of an outbreak.
Swabs used for rectal screening were placed in Amies agar gel transport medium with charcoal (Copan Italia) and were cultured on chromogenic media to screen for vancomycin-resistant Enterococcus faecium (purple colonies) and vancomycin-resistant Enterococcus faecalis (> 10 blue colonies) (Brilliance VRE Agar, Oxoid). Incubation took place at 37°C for 22-24 hours in the dark. Vancomycin resistance was confirmed by sampling the cultures, instilling the samples in Triton X-100 lysis buffer and performing polymerase chain reaction (PCR) for the vanA and vanB genes (primers developed internally). An intermediate saline dilution method was used for presumed vancomycin-resistant E. faecalis colonies. Screening results were deemed positive if both culture and PCR results were positive and were otherwise deemed negative.
When clinically indicated, Enterococcus was cultured and identified from sterile-site specimens according to routine practices for the site of origin. Possible Enterococcus was identified by the presence of colonies of gram-positive cocci in pairs and chains in appropriate culture media. Species identification was confirmed using Vitek automated biochemical detection (April 2010-August 2014) or the Vitek MS system (August 2014-December 2015) (bioMérieux). Susceptibility testing was performed using the Vitek automated biochemical system (bioMérieux).
Covariables
We extracted the following covariables from the database: age, sex, ward, service category, sterile site, Enterococcus species, and time between swab screening and sterile-site sampling for culture. We used these data to inform the baseline characteristics of screened patients.
Statistical analysis
We examined the following test characteristics of VRE screening swab results for their value in predicting vancomycin resistance in sterile-site isolates of Enterococcus: sensitivity, specificity, negative predictive value, positive predictive value, positive likelihood ratio and negative likelihood ratio. The VRE screening results and the vancomycin sensitivity of sterile-site isolates were deemed to represent the test and response variables, respectively. A positive screening result combined with a vancomycin-resistant sterile-site isolate was deemed a true positive. A positive screening result combined with a vancomycin-sensitive sterile-site isolate was deemed a false positive. A negative screening result combined with a vancomycin-sensitive sterile-site isolate was deemed a true negative. A negative screening result combined with a vancomycin-resistant sterile-site isolate was deemed a false negative. We calculated confidence intervals (CIs) using the Clopper-Pearson exact method, with the efficient score method used for negative predictive value in the "immediate" group (described below) and the log method used for CIs for likelihood ratios.21-23 All test characteristics were calculated using Stata 14.1 (Statacorp).
We performed subgroup analyses based on the species of Enterococcus isolated in culture of sterile-site specimens and the time between swab screening and sterile-site sampling. We categorized these intervals, on the basis of prior work on methicillin-resistant S. aureus,14 as immediate (0-48 h), recent (48 h-14 d) and remote (> 14 d).
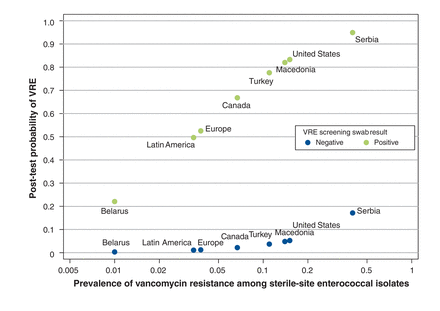
The local prevalence of vancomycin resistance among Enterococcus isolates and intrinsic test characteristics for VRE swab screening both contribute to the post-test probability of vancomycin resistance in Enterococcus isolates. We plotted post-test probability (y axis) as a function of prevalence (x axis, logarithmic) and denoted representative countries for which published VRE prevalence data were available. We derived prevalence data from published repositories of surveillance data on antimicrobial resistance for various countries and regions.24-26 We generated the plot using Stata 14.1.
We calculated number needed to screen using the total number of unique patients with VRE screening swab results divided by the number of unique patients with sterile-site specimens growing Enterococcus who had prior VRE screening.
Ethics approval
The study was approved by the Sunnybrook Health Sciences Centre Research Ethics Board.
Results
Availability of VRE screening results and prevalence of vancomycin resistance
Culture results for sterile-site specimens were positive for Enterococcus species for a total of 619 patients. Of these, 488 (79%) had prior results for VRE screening; for 347 (71%) of these 488 patients, the sterile-site culture result was positive for E. faecalis, and for 141 (29%), it was positive for E. faecium. A total of 19 (4%) of the 488 screening results were positive for VRE, and 10 (2%) of the sterile-site Enterococcus isolates were resistant to vancomycin. All VRE-positive sterile-site specimens grew E. faecium (prevalence of vancomycin-resistant E. faecium 7% [10/141]). Of the 131 patients who were excluded (because a prior screening result was not available), 3 had a positive culture result for VRE, and 2 of these 3 patients had a positive result on swab screening for VRE later the same day. Baseline patient characteristics are presented in Table 1.
Utility of prior VRE screening swab results in predicting vancomycin resistance in subsequent Enterococcus sterile-site infections
The utility of swab screening for VRE in predicting vancomycin resistance in Enterococcus sterile-site infections is summarized in Table 2. The specificity of VRE swabs was 97% (95% CI 96%-99%), and the sensitivity was 70% (95% CI 35%-93%). The post-test probability of VRE in our study population, given a positive VRE swab screening result, was 37%, equivalent to the positive predictive value of 37% (95% CI 16%-62%); the negative predictive value was 99% (95% CI 98%-100%). The positive likelihood ratio was 28 (95% CI 14-56) and the negative likelihood ratio was 0.31 (95% CI 0.12-0.79). Most patients who undergo surveillance for VRE with swab screening do not subsequently experience an enterococcal infection; in our study population, the number needed to screen was 103 to have swab information available for 1 patient with subsequent enterococcal infection.
In a post hoc sensitivity analysis limited to patients with blood culture results for Enterococcus (n = 351), the results were similar in direction and magnitude to the overall study findings (data not shown).
Subgroup analyses
In the "immediate" subgroup (interval of 0-48 h between swab screening and sterile-site sampling for culture), the specificity was 99% (95% CI 93%-100%) and the sensitivity 100% (95% CI 16%-100%). In the "recent" subgroup (interval of 48 h-14 d), the specificity was 97% (95% CI 94%-99%) and the sensitivity 67% (95% CI 9%-99%). In the "remote" subgroup (interval > 14 d), the specificity was 97% (95% CI 94%-99%) and the sensitivity 60% (95% CI 15%-95%). Of note, the positive predictive value of VRE swab screening was 67% (95% CI 9%-99%) for patients in the "immediate" subgroup, and the positive likelihood ratio was 74 (95% CI 11-518).
The overall positive predictive value for E. faecium was 64% (95% CI 31%-89%), and the positive likelihood ratio was 23 (95% CI 8.1-65). Fourteen patients had swabs taken and culture samples drawn in the "immediate" period that grew E. faecium; in this context, the swab screening was 100% sensitive (95% CI 16%-100%) and 100% specific (95% CI 74%-100%).
Post-test probability of vancomycin resistance
Figure 1 shows how the local prevalence of VRE informs the post-test probability of vancomycin resistance among sterile-site Enterococcus infections with prior VRE swab results. Specifically, small differences in prevalence lead to substantial changes in post-test probability. For example, a 15.1% prevalence of VRE in bloodstream isolates, such as that seen in a cohort of patients from North American hospitals (mostly in the United States), would translate to a post-test probability of VRE of 83% among patients with a prior positive swab result.25
The post-test probability of vancomycin-resistant Enterococcus (VRE) among patients with sterile-site Enterococcus infection and a positive or negative result on prior swab screening for VRE, as a function of regional prevalence of VRE (shown with a logarithmic scale). For each country, 2 data points are shown: the lower data point reflects post-test probability among patients with a negative swab result, and the upper data point reflects post-test probability among patients with a positive swab result. Each point represents published data on the regional prevalence of VRE among sterile-site isolates of Enterococcus24-26 and the calculated post-test probability based on the positive likelihood ratio derived in the current study.
Interpretation
We explored the utility of swab screening for VRE in predicting vancomycin resistance in subsequent Enterococcus sterile-site infections. The sensitivity of swab screening was 70%, and specificity was 97%. The positive likelihood ratio was 28, and the negative likelihood ratio was 0.31. The high specificity of VRE swab screening was preserved for each defined interval between swab screening and sterile-site sampling, whereas sensitivity diminished with increasing time. Test characteristics were optimized for E. faecium. The results show that among patients with a prior positive result on VRE swab screening and Enterococcus isolates identified in sterile-site cultures, VRE is sufficiently probable that empiric coverage is necessary. Similarly, a negative result on VRE swab screening may be sufficient to withhold empiric VRE coverage in settings of intermediate VRE prevalence.
This work shows the value of VRE swab screening in predicting resistance among Enterococcus sterile-site isolates. Rolston and colleagues19 looked at patients with hematologic malignancies and found that VRE colonization had a positive predictive value of 29.3% (29/99) for VRE bacteremia and a negative predictive value of 99.9% (2014/2016). Kim and colleagues27 described a prospective study in an intensive care unit in which 9.2% (17/184) of VRE-colonized patients experienced VRE sterile-site infections. In a study of high-risk patients with febrile neutropenia, Bossaer and coworkers18 showed a 26.4% rate (14/53) of VRE bloodstream infection in VRE-colonized patients. Finally, Ford and colleagues17 found that 12.2% (10/82) of VRE-colonized patients with leukemia later experienced a VRE bloodstream infection. All of these studies involved specialized clinical settings and immunocompromised patients. More importantly, they examined the overall risk of VRE infection in VRE swab-positive patients,16 whereas we examined how prior VRE swab results influence the conditional probability of vancomycin resistance once an Enterococcus infection is documented.
Our study informs, in a novel fashion, the empiric treatment decision in the window between identification of a pathogen and availability of susceptibility results.13 This important window has recently widened because of the availability of rapid methods for species identification.13 As with methicillin-resistant S. aureus, the current findings are mechanistically consistent with the long duration of rectal colonization with VRE and with the potential to acquire VRE carriage during the hospital stay.28 The median duration of VRE colonization has been estimated at 4 to 9 weeks in some studies.8,28,29 Our results are also consistent with rectal VRE colonization being an intermediate step in the acquisition of a VRE sterile-site infection. This theoretical basis is supported by the strong correlation in genotyping of VRE isolates from matched blood cultures and rectal swabs.16,17 It makes intuitive sense that VRE colonization, the last step before VRE infection, would provide useful predictive information for a subsequent infecting isolate.
Our analysis of regional variation in the post-test probability of VRE sterile-site infection shows the influence of VRE prevalence. Local information can be used to decide whether the probability of VRE infection is sufficient to inform starting or withholding empiric therapy. Recent work has highlighted that patient survival may be directly related to appropriate early empiric coverage of VRE.12 Once the prevalence of vancomycin resistance in Enterococcus sterile-site culture isolates increases beyond 1%, the post-test probability of VRE sterile-site culture isolates in a patient with a prior positive result on VRE swab screening exceeds 20%, which necessitates empiric coverage with agents such as linezolid or daptomycin.1,6,30 This is the most important implication of our study to clinical practice.
Moreover, when Enterococcus sepsis is of serious concern in patients colonized with VRE - for instance, patients with hematologic malignant disease and those in critical care units - we suggest coverage for VRE immediately upon suspicion of serious infection, without waiting for species identification. Finally, when the prevalence of vancomycin resistance in Enterococcus sterile-site culture isolates is 15% or lower, the post-test probability of VRE sterile-site culture isolates in a patient with a prior negative result on VRE swab screening is at most 5%, and consideration may be given to withholding empiric VRE coverage.
Limitations
The potential limitations of this study relate to its retrospective design and the consequent potential for bias. The accuracy of test characteristics depends on culture being performed for all sterile-site infections and on the accuracy of culture and screening methods. Restriction of the study to sterile-site culture results provides confidence that positive culture results reflect true infection. The current results were obtained for patients who met VRE screening criteria and may not be directly transferable to centres that employ universal screening. Although the study was conducted in a population with low baseline VRE prevalence, the intrinsic test characteristics should be broadly applicable. The wide CIs for some test characteristics reflect low VRE prevalence in our setting. We anticipate that future work performed at multiple centres and in areas of higher VRE prevalence will confirm the broad trends that we have described. Given the small number of positive events, the subgroup analysis by time between swab screening and sampling for culture can only be used to generate hypotheses for future work. Importantly, VRE screening swabs are already available and are employed for other purposes at many centres; the conclusions of our study can therefore be implemented with little delay. The number needed to screen to have a VRE screening swab result available for use in determining empiric therapy for an Enterococcus sterile-site isolate was 103 for our study population. Given the cost of VRE screening, we do not expect this novel clinical use, on its own, to justify broad screening of patients. Future work may show more limited cost-effectiveness for screening susceptible patients in high-risk settings.
Conclusion
Our study has shown that positive results on VRE swab screening reliably increase the probability of vancomycin resistance in Enterococcus sterile-site culture isolates, beyond usual thresholds meriting empiric coverage. New technologies have increased the time between species identification and availability of antimicrobial susceptibilities. Our results suggest that VRE should be covered during this empiric window for patients with prior positive results on VRE swab screening and a current sterile-site specimen growing Enterococcus.
Supplemental information
For reviewer comments and the original submission of this manuscript, please see www.cmajopen.ca/content/5/3/E632/suppl/DC1.
Footnotes
Competing interests: None declared.
Contributors: All of the authors contributed to development of the study concept and design, conduct of the analyses and interpretation of the results. All of the authors participated in drafting the manuscript and revising it for important intellectual content, gave final approval of the version to be published and agreed to be accountable for all aspects of the work.
References
- Copyright 2017, Joule Inc. or its licensors