Abstract
Background: As Canada increases requirements for research data management and sharing, there is value in identifying how research data are shared and what has been done to make them findable and reusable. This study aimed to understand Canada’s data-sharing landscape by reviewing how data funded by the Canadian Institutes of Health Research (CIHR) are shared and comparing researchers’ data-sharing practices to best practices for research data management and sharing.
Methods: We performed a descriptive analysis of CIHR-funded publications from PubMed and PubMed Central published between 1946 and Dec. 31, 2019, that indicated that the research data underlying the results of the publication were shared. We analyzed each publication to identify how and where data were shared, who shared data and what documentation was included to support data reuse.
Results: Of 4144 CIHR-funded publications identified, 1876 (45.2%) included accessible data, 935 (22.6%) stated that data were available via request or application, and 300 (7.2%) stated that data sharing was not applicable or possible; we found no evidence of data sharing in 1558 publications (37.6%). Frequent data-sharing methods included via a repository (1549 [37.4%]), within supplementary files (1048 [25.3%]) and via request or application (935 [22.6%]). Overall, 554 publications (13.4%) included documentation that would facilitate data reuse.
Interpretation: Publications funded by the CIHR largely lack the metadata, access instructions and documentation to facilitate data discovery and reuse. Without measures to address these concerns and enhanced support for researchers seeking to implement best practices for research data management and sharing, much CIHR-funded research data will remain hidden, inaccessible and unusable.
To improve health outcomes and research reproducibility, health sciences research has become increasingly focused on the production, management and sharing of research data. Increased interest in making this research more reproducible and reusable has set in motion initiatives in the United States,1,2 Europe,3,4 Canada5 and internationally6 to improve data discoverability, accessibility and transparency.
The importance of data sharing in this area is well documented. Sharing research data catalyzes new research discoveries; 7–12 encourages transparency and holds the research community accountable;13–15 and improves the interoperability of data across research communities and systems.16–18
Canada is at a crucial stage of development with respect to data-management and data-sharing initiatives. The Canadian Tri-Agency has released a policy on research data management that requires researchers to manage and deposit data,19 Canadian publishers are releasing data-sharing policies,20 the Federated Research Data Repository21 has made it possible to discover data that are produced and stored in Canadian repositories, and the New Digital Research Infrastructure Organization has been established to respond to emerging data needs within the Canadian digital research landscape.22 Although these efforts aim to make data sets more discoverable, valuable data shared alongside publications, in external discipline-specific repositories, via websites or on request are difficult to locate, access and reuse.
With the release of the Tri-Agency Research Data Management Policy19 and the establishment of new initiatives to locate Canadian research products online, we see value in identifying how and where Canadian research data are being shared, and what steps have been taken to make them reusable. In the present study, we aimed to understand the Canadian data-sharing landscape by reviewing how and where data funded by the Canadian Institutes of Health Research (CIHR) are shared, and comparing the data-sharing practices of CIHR-funded researchers to the Tri-Agency principles for research data management and sharing.23
Methods
Design and setting
We used descriptive analysis to identify how and where CIHR-funded researchers share their data, using metadata extracted from CIHR-funded publications. The CIHR is Canada’s federal funding agency for health research. For the purpose of this study, metadata are defined as the data used to describe specific elements (e.g., authors, title, provenance) of a publication, data set or research product to make it searchable and interpretable.24 This study began in October 2019 and was completed in October 2020.
Data sources
We searched PubMed and PubMed Central databases to identify CIHR-funded publications indicating that they shared research data underlying the published results. PubMed is a freely available bibliographic database that contains more than 28 million citations of biomedical literature from the MEDLINE database, life sciences journals and online books. PubMed Central is a freely available digital repository that archives open-access full-text scholarly publications in biomedical and life sciences journals. Reciprocal links exist between the full text in PubMed Central and the corresponding citations in PubMed.25 We did not search for data associated with the grey literature because our study focused on CIHR-funded scholarly publications.
We chose PubMed and PubMed Central as our data sources because they provide unique data set search filters26 that identify publications indicating that data underlying the results have been shared. We define research data as
data that are used as primary sources to support technical or scientific enquiry, research, scholarship, or artistic activity, and that are used as evidence in the research process and/or are commonly accepted in the research community as necessary to validate research findings and results.27
Search strategy
Our search strategy, which was built by K.B.R. and peer reviewed by H.G. and D.R.S., identified publications between 1946 and Dec. 31, 2019. Using PubMed Central’s data filters, 26 1 of the authors (K.B.R.) searched for all CIHR-funded publications that included a statement on data availability. Data-availability statements contain the authors’ description of where and how to gain access to the research data underlying the published manuscript. K.B.R. searched for additional publications in PubMed using its data filter,26 which captures when data have been shared in a repository. One author (K.B.R.) then combined these filters with CIHR-related keywords in English and French, using the grants information field from both databases (Appendix 1, available at www.cmajopen.ca/content/9/4/E980/suppl/DC1).
Publications were included if, in both PubMed and PubMed Central, they had an author who was funded by the CIHR, were published between 1946 and Dec. 31, 2019, and were published in English or French (the 2 official languages of CIHR-funded research). Publications from PubMed were required to link to a data repository, and publications from PubMed Central were required to include a data-availability statement.
Publications were excluded if, in both PubMed and PubMed Central, they did not include a CIHR-funded author and if, in PubMed, they linked exclusively to a clinical trial registry rather than a data repository.
Metadata extraction
One author (K.B.R.) extracted selected metadata fields from publications that met our inclusion criteria using the PubMed Central Open Access Subset,28 which allows full-text metadata from a publication to be extracted under a Creative Commons licence. When full-text metadata were not available via the Open Access Subset, they were extracted with the use of the minimal level of metadata available in PubMed Central. Publications that were not available in PubMed Central had a limited set of metadata extracted from PubMed (Appendix 2, available at www.cmajopen.ca/content/9/4/E980/suppl/DC1). The Python scripts used to extract the metadata from both databases are available via the Open Science Framework.29
Data abstraction
Examination of data-sharing practices
Using the extracted metadata, we examined each publication to explore data accessibility; how, where and by whom research data were shared; and the inclusion of documentation to support data reuse, including but not limited to codebooks, data-analysis plans, software code and readme files. Each author was assigned a unique set of publications to examine independently. Any uncertainties that arose during the data-collection process were resolved at biweekly team meetings. Our data-collection instrument and descriptive analyses were generated and captured in a REDCap database. The instrument and data dictionary used for our final analysis are available via the Open Science Framework.29
Before starting the complete examination of all publications, each author examined a random preliminary set of the same 150 publications to ensure consistency in how information was interpreted and captured in our data-collection instrument.
Data-sharing status
To frame our analysis, we grouped data-sharing practices into categories representing the most commonly identified data-sharing status types (Table 1); these categories are not mutually exclusive. We examined the frequency of each category across our entire sample and over time.
Data-sharing status categories and their definitions
Data-sharing methods
Using the metadata available (Appendix 2) and building on our data-sharing status categories, we recorded the methods of data sharing evident within each publication. These included but were not limited to sharing data via a repository, within the supplementary files, via request or application, within the publication, via a website or when an author stated that data sharing was not applicable or possible.
If an author’s data-sharing statement indicated that an application was required to access the data, we captured all reasons why authors insisted on this requirement. Similarly, if an author stated that data could not be shared at all, we captured all reasons provided why this was the case.
We examined whether data-sharing statements made by authors aligned with how data were shared in practice. When authors stated that all the research data needed to understand the results were within the publication, we reviewed the publication for evidence that no additional research data files were needed to understand the findings. When authors stated that research data were available in the supplementary files, we attempted to locate and access the data within the supplementary files section. We documented instances of misalignment between author statements and whether and how data were shared, as well as when we were unclear about whether author statements reflected data sharing accurately.
Finally, we categorized institutions and journals where data were shared and ranked them according to their data-sharing status.
Research data documentation
We captured the types of documentation that were included alongside accessible and available research data (Table 1, categories 1 and 2). We identified types of documentation based on the Tri-Agency Statement of Principles on Digital Data Management,23 which makes recommendations on adherence to standards, data collection and storage, and metadata documentation. We then examined each publication to determine whether documentation such as study protocols, data-analysis plans, software code, data dictionaries, readme files, data-collection instruments, videos and data-management plans was provided. Documentation of this kind has been identified as necessary for improving the transparency, reproducibility and reusability of research results.30–33 We also analyzed the frequency of documentation inclusion over time.
Statistical analysis
We performed a descriptive analysis of our results. All data collected during this study were exported from the REDCap database and analyzed with Stata/SE 16.0 software (Stata-Corp). The raw data extracted from PubMed and PubMed Central, the synthesized data exported from REDCap and the analyzed data from Stata along with a summary analysis report are available via the Open Science Framework.29
Ethics approval
Because our study focused on a descriptive analysis of metadata that are publicly available, ethics approval was not required.
Results
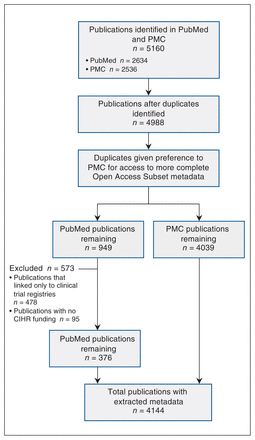
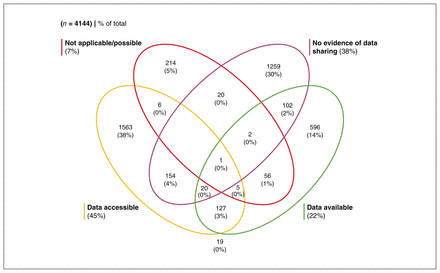
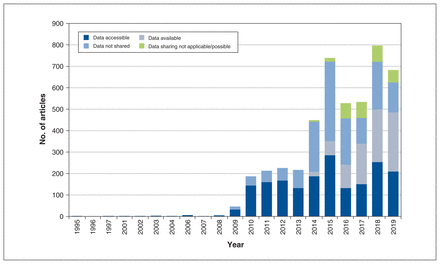
Our search identified 4988 publications. After we applied our inclusion criteria, we retrieved metadata for 4144 publications for analysis (Figure 1). Of these publications, 1876 (45.3%) made their data accessible, 935 (22.6%) made their data available (via request or application), 300 (7.2%) indicated that data sharing was not applicable or possible, and 1558 (37.6%) provided no evidence of data sharing even though we had limited our sample to publications that had indicated data sharing of some kind. Many publications shared multiple data sets in different ways. Figure 2 shows the extent of overlap among the categories of data-sharing status, and Figure 3 shows the frequency of these categories over time.
Flow diagram showing metadata extraction. Note: CIHR = Canadian Institutes of Health Research, PMC = PubMed Central.
Frequency of publications by data-sharing status.
Frequency of data-sharing status categories over time. Categories are not mutually exclusive.
Data-sharing methods
The most frequent methods of data sharing were via a repository (1549 publications [37.4%]) and within the supplementary files (1048 [25.3%]) (Table 2). Notably, 935 publications (22.6%) stated that data were available via request (701 [16.9%]) or application (234 [5.6%]) but provided little detail about how to acquire the data. A total of 538 publications (13.0%) had no evidence of or information about data sharing whatsoever. Some publications shared data in multiple formats and therefore may be represented in more than 1 category.
Frequency of data-sharing methods
Among the 1549 publications that reported data sharing via a repository, there were 97 repositories represented (the complete listing is available in the Open Science Framework analysis report29). The most prevalent repositories were the Protein Data Bank (599 [38.7%]), Gene Expression Omnibus (377 [24.3%]) and GenBank (194 [12.5%]). A breakdown of repositories is shown in Appendix 3, Supplementary Figure S1 (available at www.cmajopen.ca/content/9/4/E980/suppl/DC1).
A total of 234 publications indicated that an application was required to access the data underlying the results. The most frequent justification for this requirement was the need to complete a data-access, data-transfer or data-use agreement (78 [33.3%]), followed by general ethics concerns (66 [28.2%]), confidentiality (60 [25.6%]), licence restrictions (28 [12.0%]) and Indigenous considerations (6 [2.6%]). Twenty-three publications (9.8%) did not explain why an application was required. None of the publications that required an application included metadata sufficiently outlining the requirements for access and approval.
Among the 300 publications that indicated that data sharing was not applicable or possible, the most common reason cited was confidentiality (109 [36.3%]); 88 publications (29.3%) provided no justification at all (Table 3).
Reasons for not sharing data
Of the 1048 publications stating that data were available in the supplementary files, 752 (71.8%) did not share data in this way. Similarly, 345 (39.7%) of the 870 publications stating that all data were available within the article shared no research data underlying the results within the publication or supplementary files, although there was clear evidence of data collection.
A breakdown of institutions associated with the publications that shared data is presented in Appendix 3, Supplementary Figure S2. Among institutions associated with more than 10 publications, those with the greatest proportion of publications in which data were accessible or available were the Structural Genomics Consortium (20/21 [95%]) and the University of Waterloo (12/25 [48%]), respectively.
The journals used most frequently were PLoS One (736 [17.8%]) and the Journal of Biological Chemistry (208 [5.0%]) (Appendix 3, Supplementary Figure S3A). Among journals with more than 25 CIHR-funded publications, the 3 most commonly used journals that included examples of accessible data were the Journal of Molecular Biology (57/59 [96.6%]), Proceedings of the National Academy of Sciences of the United States of America (110/116 [94.8%]) and Nature (51/54 [94.4%]). Among all journals, the 3 most commonly used that included examples of available data were the International Journal of Behavioural Nutrition and Physical Activity (23/26 [88.5%]), BMC Psychiatry (13/17 [76.5%]) and BMC Medical Research Methodology (17/23 [73.9%]) (Appendix 3, Supplementary Figure S3B and available on the Open Science Framework29).
Research data documentation
The documentation provided alongside publications was varied, with supplementary figures or tables or both, study protocols, research data files and transparent reporting forms most frequently represented (Table 4).
Documentation identified, by data-sharing status category
Referring to the recommended documentation types outlined in the Tri-Agency Principles on Digital Data Management, 23 we examined how frequently these were included alongside publications that made data accessible or indicated that data were available over time (Appendix 3, Supplementary Figure S4). The types of documentation required to understand and reuse research data were provided in a minority of publications that shared data (554/4144 [13.4%]). Across all publications regardless of whether they shared data, study protocols were most frequently included (576/4144 [13.9%]), and data-management plans were the least frequently included (4/4144 [0.1%]). Although documentation supporting reuse was scarce, as of 2017, there was increasing availability of data-analysis plans, code and data-collection instruments.
Interpretation
This study highlights substantial room for growth in improving the discoverability, accessibility and usability of CIHR-funded research data. Although, encouragingly, repositories were the most common venues authors chose to share data, the remaining data were made available within the publication or its supplementary files, by request or by a long tail of other means.29 When authors indicated that data were available via request or application (22.6% of publications), they did not provide adequate instructions on how to acquire them. Despite our focus on publications indicating that data had been shared, more than one-third (37.6%) showed no evidence of sharing. The most frequent types of documentation shared alongside data did not generally support their interpretation and reuse. These characteristics conflict with expectations outlined in Canada’s Tri-Agency data management23 and international FAIR (Findable, Accessible, Interoperable, Reusable) guiding principles.16
Deficiencies in data discoverability, access and usability have been examined in other contexts. A 2015 study of data sharing in publications funded by the US National Institutes of Health showed that 88% of research data were not discoverable. 34 Our finding that there is often a gap between data-sharing statements and practices is in keeping with recent studies.35–38 We identified several cases in which authors indicated that data were shared but we found no evidence thereof. We speculate that those authors may have incorrectly considered summary tables and figures to be research data. This finding highlights that many authors may fundamentally misunderstand what it means to make research data discoverable and accessible. Our analysis of reusability practices showed that the most frequent types of documentation shared alongside data rarely support their interpretation and reuse. Sharing descriptive documentation such as codebooks and data dictionaries, and actionable supporting files such as code and software, is increasingly recognized as best practice,30,33,39 and our results indicate that CIHR-funded data sharing can vastly improve in this area.
Inadequate metadata are a recognized problem in the data-sharing landscape,40–42 and the examples of CIHR-funded sharing we encountered are no different. Without adequate metadata to support discovery, data will remain hidden.43 The absence of metadata elaborating on application requirements specifically calls into question the true availability of these data and impedes future research based on them. Other investigators have highlighted the challenges of requesting access to data in relation to the lack of transparency of request processes and a lack of standardization in use agreements for health data.41,44,45 Given that data made available by request are often collected from human participants, improving the discoverability of and access to sensitive data can prevent study replication, create opportunities for pooling related data and increase research efficiency.41,42,44,45 Our findings indicate that current metadata practices do not provide sufficient information to make successful data requests and secure these outcomes.
Future initiatives should focus on the development of metadata standards that facilitate the discovery of sensitive data and support transparent data-request processes. In particular, we suggest that the Tri-Agency’s requirements for data-management plans19 be extended to include reporting guidelines for making sensitive research data discoverable. These guidelines should require robust descriptions of sensitive data and detailed data-access procedures where applicable, which could be submitted alongside a manuscript for publication. We also suggest that Canadian data repositories explore how to better accommodate sensitive data so they can be made discoverable while honouring access and privacy restrictions. We see value in reestablishing infrastructure that tracks CIHR-funded publications and data sharing, a function that was partially fulfilled by PubMed Central Canada before it was taken offline, in 2018.46
Limitations
Our analysis focused solely on CIHR-funded publications indicating that data were shared. We did not deem it logistically possible to identify instances of data sharing from the full body of CIHR-funded literature. We also examined CIHR-funded publications exclusively from PubMed and PubMed Central. Although these are the most comprehensive biomedical databases available, we acknowledge there are other databases where CIHR-funded publications exist. We limited our study to these sources because metadata specific to data sharing were not readily accessible in other biomedical databases, whereas PubMed and PubMed Central provide access to open-source metadata. To manage study feasibility, we limited our review of documentation to that which was shared or stated within the publication and did not extend this analysis to repositories or websites where some research data were shared.
Conclusion
Publications funded by the CIHR largely lack the metadata, access instructions and documentation to facilitate data discovery and reuse. Without measures to address these concerns and enhanced support for researchers seeking to implement best practices for research data management and sharing, much CIHR-funded research data will remain hidden, inaccessible and unusable.
Acknowledgement
The authors thank the Canadian Hub for Applied and Social Research for its support in procuring and analyzing the research data in this study.
Footnotes
Competing interests: None declared.
This article has been peer reviewed.
Contributors: Kevin Read supervised the project and conceived the study. All of the authors designed the study, obtained, analyzed and interpreted the data, drafted the manuscript and revised it critically for important intellectual content, approved the final version to be published and agreed to be accountable for all aspects of the work.
Funding: This project was supported in part by the University of Saskatchewan Faculty Recruitment and Retention Program.
Data sharing: All raw, processed and analyzed data, as well as accompanying documentation, reports and scripts are available on the Open Science Framework, at https://osf.io/n9jv5/.
Supplemental information: For reviewer comments and the original submission of this manuscript, please see www.cmajopen.ca/content/9/4/E980/suppl/DC1.
This is an Open Access article distributed in accordance with the terms of the Creative Commons Attribution (CC BY-NC-ND 4.0) licence, which permits use, distribution and reproduction in any medium, provided that the original publication is properly cited, the use is noncommercial (i.e., research or educational use), and no modifications or adaptations are made. See: https://creativecommons.org/licenses/by-nc-nd/4.0/
References
- © 2021 CMA Joule Inc. or its licensors